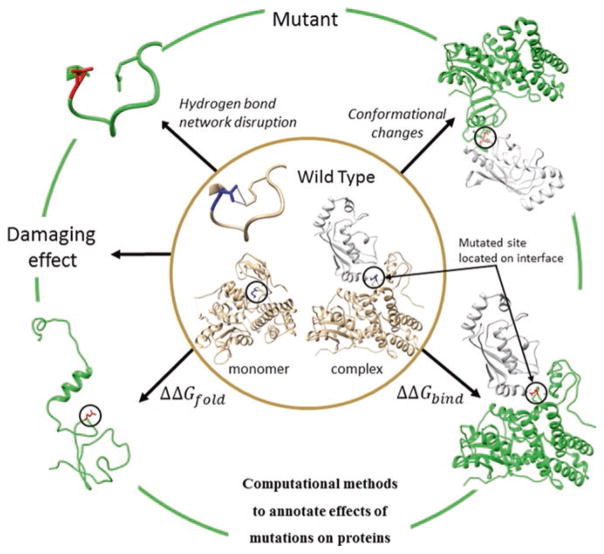
Molecular Mechanisms of Disease Mutations
Mutations can render proteins dysfunctional and may be responsible for many diseases. However, the molecular mechanisms of mutation effects on proteins remain unclear. We develop computational approaches to make structural models of mutated proteins and estimate the quantitative effects of mutations on protein interaction networks, binding and pathways. Our methods are based on using molecular mechanics force fields, statistical potentials, fast side-chain optimization and statistical algorithms. This help to discern the driver effects of mutations.